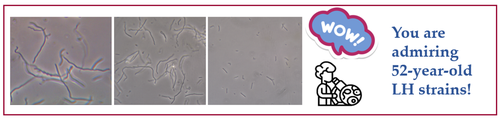
This project aimed to investigate the evolutionary dynamics of Lactobacillus helveticus (LH) in traditional dairy environments over the past fifty years. The work was based on two extensive LH collections: one isolated and preserved in 1970 from natural whey starters (NWS) used for PDO cheese production, never revitalized, and deposited at DISTAL (UNIBO) and a second collection isolated in 2021–2023 from the same ecological and technological context. The availability of these historical and contemporary collections provided a unique opportunity to compare LH biodiversity within the same production environment.
The strains were initially identified and genotypically characterized by AFLP analysis, which revealed high biodiversity and a clear differentiation between historical and recent isolates. Based on preliminary genotypic and phenotypic screening, 16 representative strains (including both old and new isolates) were selected for further physiological, technological, and whole-genome sequencing analyses.
Comparative genomic analysis highlighted evolutionary changes over time, suggesting a possible genome reduction phenomenon in contemporary strains. From a phenotypic perspective, historical strains showed higher growth rates and shorter lag phases in culture medium and milk. In miniaturized cheese models simulating cooked hard cheese production, differences emerged in microbial dynamics and metabolite production: historical strains tended to decrease more rapidly in cell concentration, likely due to lysis, accumulated lower amounts of acetic acid and acetoin, but produced higher levels of alcohols during ripening.
Based on genomic, physiological, and technological results, four strains were selected for pilot-scale cheesemaking trials: two historical and two recent. The strains were co-inoculated with a Lactobacillus delbrueckii subsp. lactis UPCCO strain to reproduce cultures similar to traditional natural whey starters. Experimental cheeses (~1 kg) were produced in biological duplicates using pasteurized milk and a protocol simulating cooked hard cheese production, followed by ripening at 16 °C for three weeks.
Microbial dynamics were monitored through plate counts and qPCR, while volatile compound profiles were analyzed by SPME-GC-MS. The results highlighted functional and metabolic differences between historical and recent strains in terms of microbial development and volatile compound production, providing new insights into microbial evolution within dairy fermentation ecosystems and supporting the targeted selection of strains for specific dairy applications. The study underscored the remarkable importance of preserving historical microbial biodiversity as a biotechnological resource for the selection of new strains of industrial interest.